obliqueslice
Extract oblique slice from 3-D volumetric data
Syntax
Description
B = obliqueslice(V,point,normal)V. The slice is
extracted with reference to a given point on the volume and a normal vector. The slicing
plane is perpendicular to the normal vector and passes through the specified point.
For information about how the slice is extracted with respect to the given point and the normal, see Oblique Slicing.
Examples
Load a 3-D volumetric data set into the workspace.
load mriRemove singleton dimensions by using the squeeze function.
V = squeeze(D);
Display horizontal slices of the data by using the montage function.
montage(V,map,'Size',[3 9]);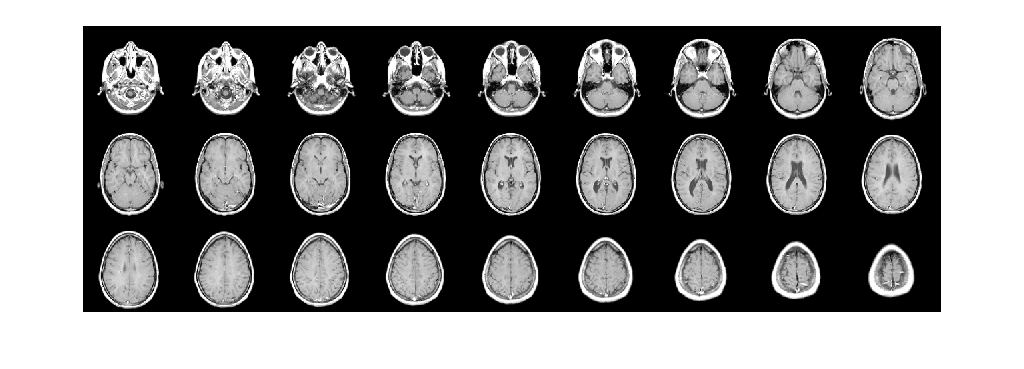
Specify a point in the volume for the slice to pass through.
point = [73 50 15.5];
Specify a normal vector in 3-D coordinate space.
normal = [0 15 20];
Extract a slice from the volumetric data. The slice is perpendicular to the normal vector and passes through the specified point.
[B,x,y,z] = obliqueslice(V,point,normal);
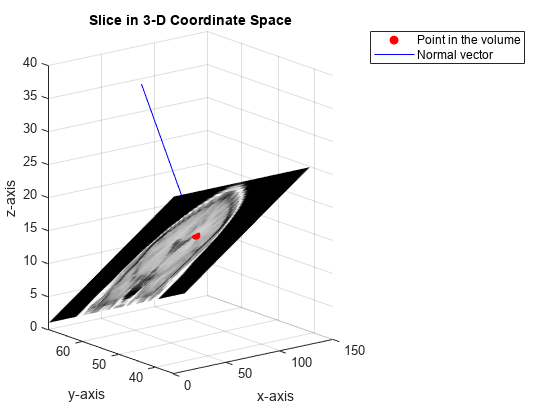
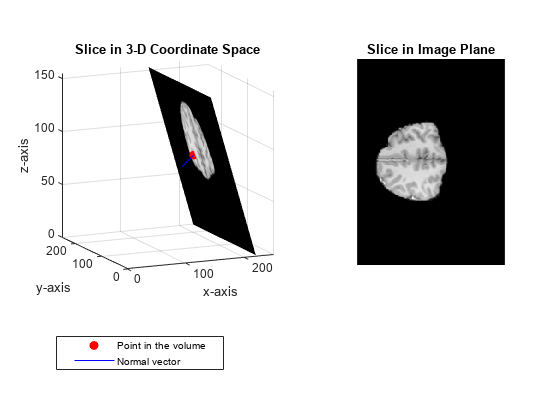
Display the extracted slice in the 3-D coordinate space.
figure surf(x,y,z,B,'EdgeColor','None','HandleVisibility','off'); grid on view([-38 12]) colormap(gray) xlabel('x-axis') ylabel('y-axis'); zlabel('z-axis'); title('Slice in 3-D Coordinate Space')
Plot the point and the normal vector.
hold on plot3(point(1),point(2),point(3),'or','MarkerFaceColor','r'); plot3(point(1)+[0 normal(1)],point(2)+[0 normal(2)],point(3)+[0 normal(3)], ... '-b','MarkerFaceColor','b'); hold off legend('Point in the volume','Normal vector')
Display the extracted slice in the image plane.
figure
imshow(B,[])
title('Slice in Image Plane')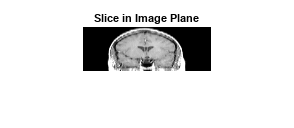
Load a 3-D volumetric data set into the workspace.
s = load(fullfile(toolboxdir('images'),'imdata','BrainMRILabeled','images','vol_001.mat')); V = s.vol;
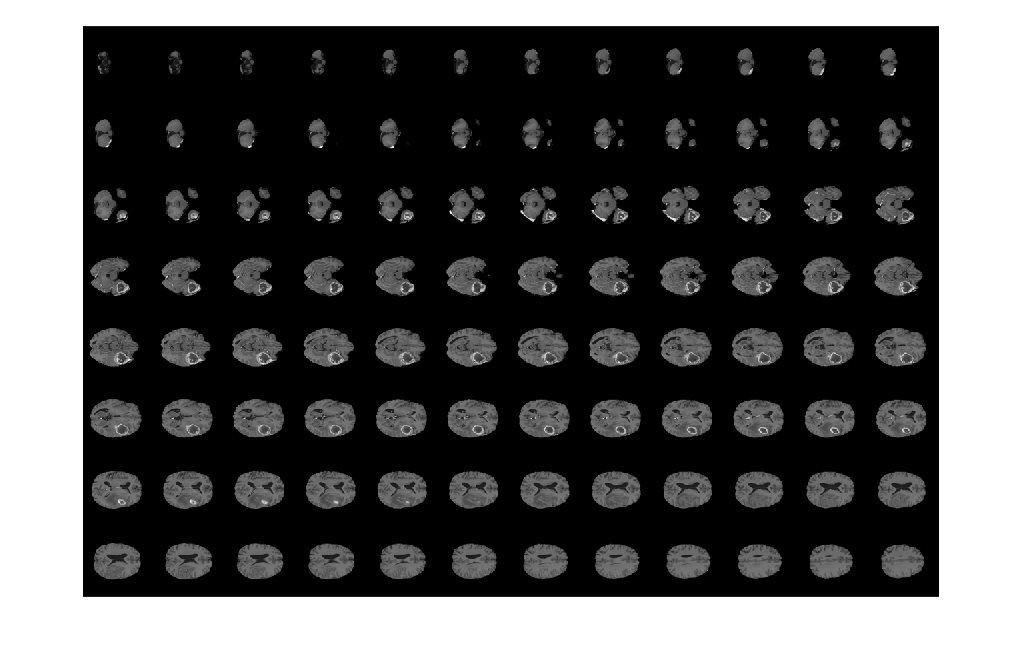
Display horizontal slices of the data by using the montage function.
montage(V,'Indices',12:118,'Size',[8 12],'DisplayRange',[]);
Specify the normal vector to a plane in 3-D coordinate space.
normal = [20 0 10];
Extract multiple slices along the direction of the normal vector using a for loop. In each iteration:
Specify a point that the slice has to pass through.
Extract the slice, specifying the output size to
'Full'and the fill value for padding pixels as 255. The extracted slices are perpendicular to the normal vector and pass through the specified point.Display the extracted slices.
sliceIdx = 10:5:180; figure for s = 1:length(sliceIdx) pt = [sliceIdx(s) 150 80]; [B,x,y,z] = obliqueslice(V,pt,normal,'OutputSize','Full','FillValues',255); Bslices(:,:,s) = B; % Display the slice in 3-D coordinate space subplot('Position',[0.11 0.36 0.38 0.5]) surf(x,y,z,B,'EdgeColor','None','HandleVisibility','off'); grid on view([-24 12]) colormap(gray) xlabel('x-axis') ylabel('y-axis'); zlabel('z-axis'); zlim([0 155]); ylim([0 250]); xlim([0 250]); title('Slice in 3-D Coordinate Space') % Plot the point and the normal vector. hold on plot3(pt(1),pt(2),pt(3),'or','MarkerFaceColor','r') plot3( ... pt(1)+[-normal(1) normal(1)], ... pt(2)+[-normal(2) normal(2)], ... pt(3)+[-normal(3) normal(3)], ... '-b','MarkerFaceColor','b') legend('Point in the volume','Normal vector','Position',[0.1 0.12 0.3 0.08]) hold off % Display the extracted slice. subplot('Position',[0.6 0.37 0.34 0.49]) imshow(B,[]) title('Slice in Image Plane') pause(0.5); end

Display the extracted image slices by using the montage function.
figure montage(Bslices,'Size',[5 7],'DisplayRange',[]);
Input Arguments
Input volume, specified as a 3-D numeric array or 3-D categorical array.
Data Types: single | double | int8 | int16 | int32 | uint8 | uint16 | uint32 | logical | categorical
Point in the volume, specified as a 3-element row vector of the form [px py pz].
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
Normal vector, specified as a 3-element row vector of form [a
b
c]. The normal vector is a vector that is perpendicular to a
surface or plane.
To extract an orthogonal slice, you can set the normal vector to one of these values:
[1 0 0]— Extract slice in the yz-plane.[0 1 0]— Extract slice in the xz-plane.[0 0 1]— Extract slice in the xy-plane.
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: obliqueslice(V,point,normal,OutputSize="Full")
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: obliqueslice(V,point,normal,"OutputSize","Full")
Interpolation method, specified as one of these values:
"linear"— linear interpolation"nearest"— nearest neighbor interpolation
If V is numeric, the interpolation method defaults to
"linear" but can also be specified as
"nearest". If V is categorical, then the
interpolation method must be "nearest".
Data Types: char | string
Size of output image, specified as one of these values:
"limit"— The size of the output image is the actual size of the 2-D slice with respect to the dimensions of input volume. If the extracted slice region is not square or rectangular, the function automatically pads the extracted slice region with extra pixels to yield a square or rectangular image."full"— The size of the output image may not be equal to the actual size of the 2-D slice. The size of the output image is set to the maximal slice size that can be obtained from the input volume with respect to the normal vectornormal. To resize the image, the border of the extracted 2-D slice is padded with extra rows and columns.The fill value for the padded pixels is
0by default. You can use theFillValuesname-value argument to change the value.
Data Types: char | string
Fill value for padded pixels, specified as a numeric scalar, character vector, or
missing.
When V is a numeric array, specify
0for zero padding.a numeric scalar for constant padding.
When V is a categorical array, specify
a character vector that denotes a category in the input data. To know the categories, use the
categoriesfunction.missing, if the category in input data is equal to<undefined>.
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64 | char
Output Arguments
Output 2-D slice, returned as a numeric or categorical matrix. The data type of the output slice is same as the data type of the input volume.
Data Types: single | double | int8 | int16 | int32 | uint8 | uint16 | uint32 | logical | categorical
x-coordinates of the output slice in the 3-D volume, returned as
a numeric matrix of the size same as the output slice, B.
Data Types: single
y-coordinates of the output slice in the 3-D volume, returned as
a numeric matrix of the size same as the output slice, B.
Data Types: single
z-coordinates of the output slice in the 3-D volume, returned as
a numeric matrix of the size same as the output slice, B.
Data Types: single
More About
Given a point (px,
py,
pz) and the normal vector (a, b,
c), the obliqueslice function solves the plane equation a(x-px)+b(y-py)+c(z-pz)
= 0
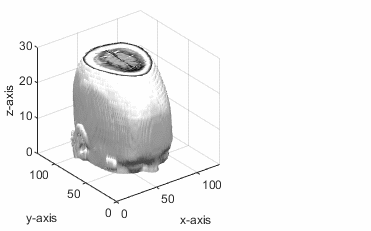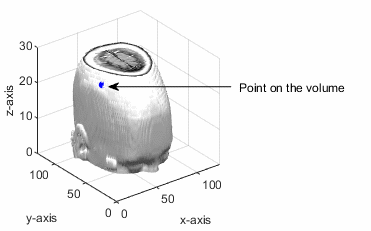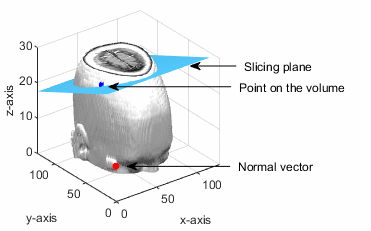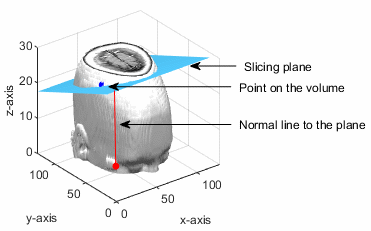
The point (px, py, pz) lies in the volumetric data. The slicing plane is perpendicular to the normal vector and passes through the given point.
The order in which the coordinates of the extracted slice in 3-D space is mapped to a 2-D plane depends on its inclination angle with respect to the horizontal and vertical planes.
The obliqueslice function returns the output
matrices x, y, and z that
contain the x, y, z coordinates of
the points in 3-D coordinate space that form the image slice. The
obliqueslice function interpolates the intensity values at these
points and maps it to the 2-D plane. The first value in the output matrices x(1,1), y(1,1),
z(1,1) specify 3-D coordinate of a point that maps as the upper-left pixel, (1,
1) in the image plane. Starting from this point, the 3-D coordinates that constitute the
image slice along with the associated intensity values are read in left-to-right,
top-to-bottom scan order. These intensity values fill the 2-D image plane in the same
left-to-right, top-to-bottom scan order.
Suppose V is the input volumetric data and B is the output 2-D image, then B(i,j) = V(a,b,c).
Where, a = x(i,j), b = y(i,j), and c = z(i,j).

Extended Capabilities
The obliqueslice function fully supports
thread-based environments. For more information, see Run MATLAB Functions in Thread-Based Environment.
Version History
Introduced in R2020aobliqueslice now supports thread-based
environments.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)