Train Classifier Using Hyperparameter Optimization in Classification Learner App
This example shows how to tune hyperparameters of a classification support vector machine (SVM) model by using hyperparameter optimization in the Classification Learner app. Compare the test data set performance of the trained optimizable SVM to that of the best-performing preset SVM model.
In the MATLAB® Command Window, load the
ionospheredata set, and create a table containing the data.load ionosphere tbl = array2table(X);Open Classification Learner using the
tbldata andYas the response variable.classificationLearner(tbl,Y)
In the Test section of the New Session from Arguments dialog box, specify to set aside
15percent of the imported data as a test data set.To accept the options and continue, click Start Session.
Train all preset SVM models. On the Learn tab, in the Models section, click the arrow to open the gallery. In the Support Vector Machines group, click All SVMs. In the Train section, click Train All and select Train All. The app trains one of each SVM model type, as well as the default fine tree model, and displays the models in the Models pane.
Note
If you have Parallel Computing Toolbox™, then the Use Parallel button is selected by default. After you click Train All and select Train All or Train Selected, the app opens a parallel pool of workers. During this time, you cannot interact with the software. After the pool opens, you can continue to interact with the app while models train in parallel.
If you do not have Parallel Computing Toolbox, then the Use Background Training check box in the Train All menu is selected by default. After you select an option to train models, the app opens a background pool. After the pool opens, you can continue to interact with the app while models train in the background.
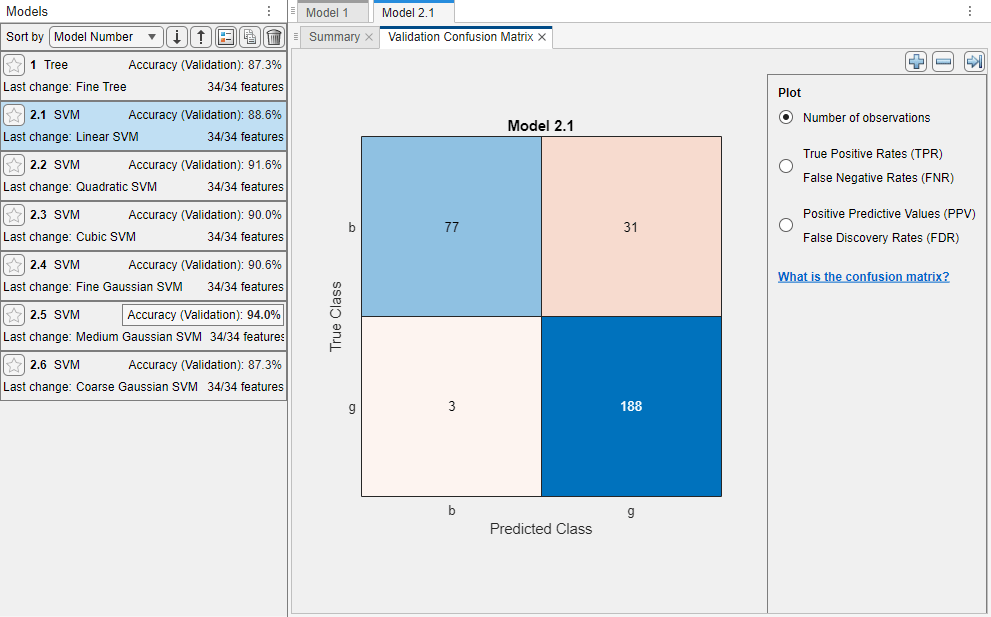
The app displays a validation confusion matrix for the first SVM model (model 2.1). Blue values indicate correct classifications, and red values indicate incorrect classifications. The Models pane on the left shows the validation accuracy for each model.
Note
Validation introduces some randomness into the results. Your model validation results can vary from the results shown in this example.
Select an optimizable SVM model to train. On the Learn tab, in the Models section, click the arrow to open the gallery. In the Support Vector Machines group, click Optimizable SVM.
Select the model hyperparameters to optimize. In the Summary tab, you can select Optimize check boxes for the hyperparameters that you want to optimize. By default, all the check boxes for the available hyperparameters are selected. For this example, clear the Optimize check boxes for Kernel function and Standardize data. By default, the app disables the Optimize check box for Kernel scale whenever the kernel function has a fixed value other than
Gaussian. Select aGaussiankernel function, and select the Optimize check box for Kernel scale.
Train the optimizable model. In the Train section of the Learn tab, click Train All and select Train Selected.
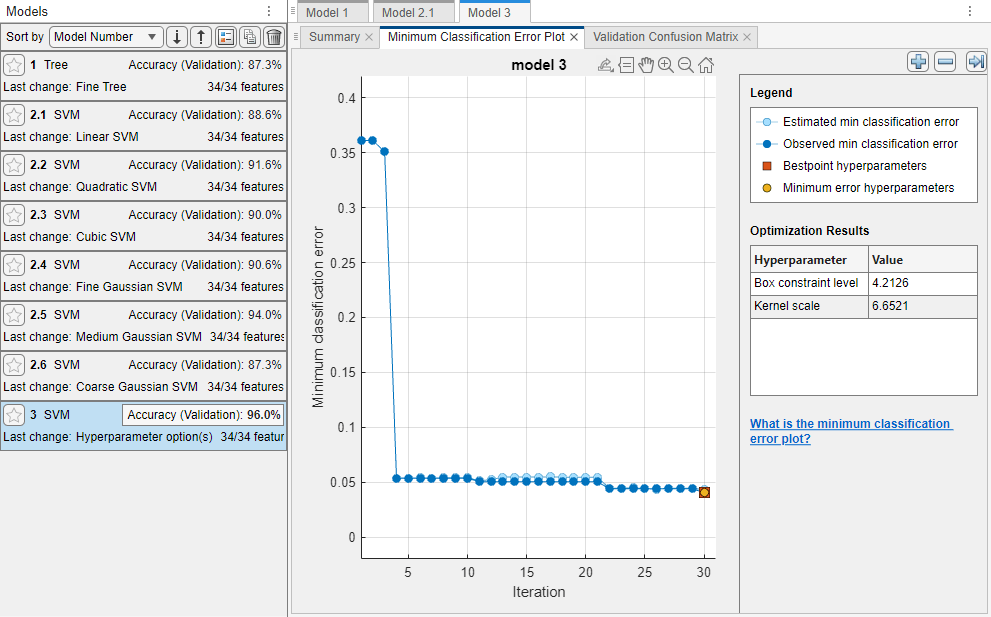
The app displays a Minimum Classification Error Plot as it runs the optimization process. At each iteration, the app tries a different combination of hyperparameter values and updates the plot with the minimum validation classification error observed up to that iteration, indicated in dark blue. When the app completes the optimization process, it selects the set of optimized hyperparameters, indicated by a red square. For more information, see Minimum Classification Error Plot.
The app lists the optimized hyperparameters in both the Optimization Results section to the right of the plot and the Optimizable SVM Model Hyperparameters section of the model Summary tab.
Note
In general, the optimization results are not reproducible.
Compare the trained preset SVM models to the trained optimizable model. In the Models pane, the app highlights the highest Accuracy (Validation) by outlining it in a box. In this example, the trained optimizable SVM model outperforms the six preset models.
A trained optimizable model does not always have a higher accuracy than the trained preset models. If a trained optimizable model does not perform well, you can try to get better results by running the optimization for longer. On the Learn tab, in the Options section, click Optimizer. In the dialog box, increase the Iterations value. For example, you can double-click the default value of
30and enter a value of60. Then click Save and Apply. The options will be applied to future optimizable models created using the Models gallery.Because hyperparameter tuning often leads to overfitted models, check the performance of the optimizable SVM model on a test data set and compare it to the performance of the best preset SVM model. Use the data you reserved for testing when you imported data into the app.
First, in the Models pane, click the star icons next to the Medium Gaussian SVM model and the Optimizable SVM model.
For each model, select the model in the Models pane. In the Test section of the Test tab, click Test Selected. The app computes the test data set performance of the model (which was trained on the training data set).
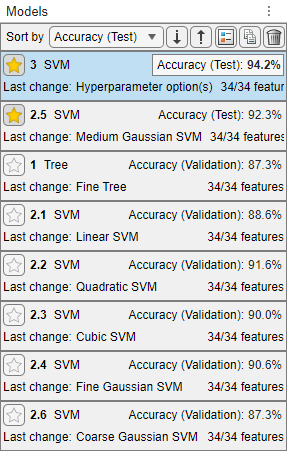
Sort the models based on the test data set accuracy. In the Models pane, open the Sort by list and select
Accuracy (Test).In this example, the trained optimizable model still outperforms the trained preset model on the test data set. However, neither model has a test accuracy as high as its validation accuracy.
Visually compare the test data set performance of the models. For each of the starred models, select the model in the Models pane. On the Test tab, in the Plots and Results section, click Confusion Matrix (Test).
Rearrange the layout of the plots to better compare them. First, close the plot and summary tabs for all models except Model 2.5 and Model 3. Then, in the Plots and Results section, click the Layout button and select Compare models. Click the Hide plot options button
 at the top right of the plots to make more
room for the plots.
at the top right of the plots to make more
room for the plots.
To return to the original layout, you can click the Layout button and select Single model (Default).
See Also
Topics
- Hyperparameter Optimization in Classification Learner App
- Train Classification Models in Classification Learner App
- Start a Classification Learner or Regression Learner Session
- Choose Classifier Options in Classification Learner
- Visualize and Assess Classifier Performance in Classification Learner
- Export Classification Model to Predict New Data
- Bayesian Optimization Workflow