cvloss
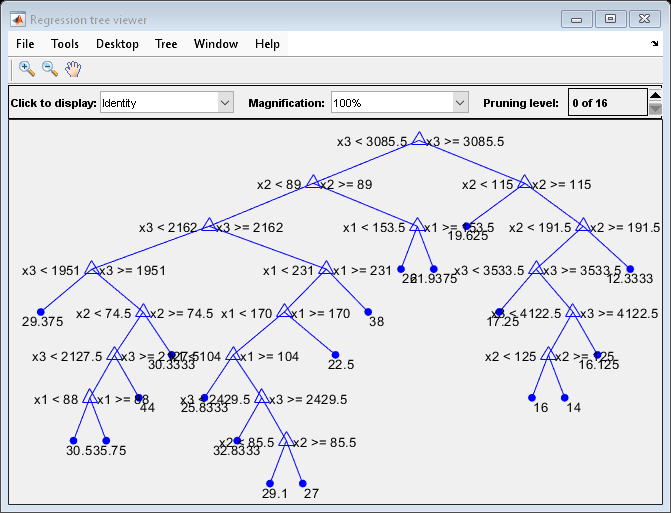
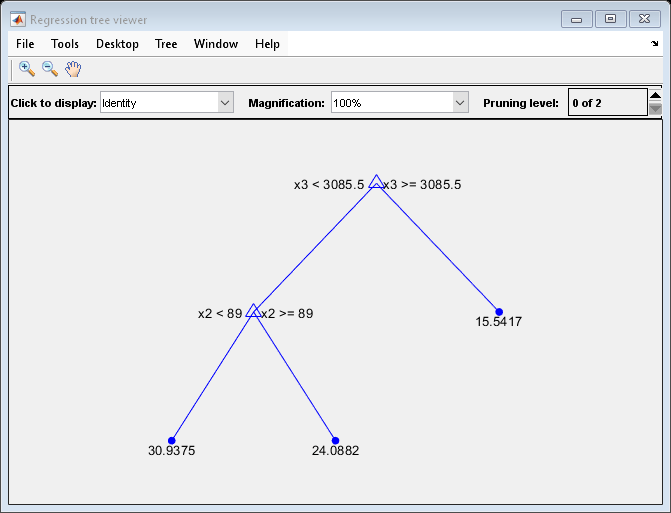
Regression error by cross-validation for regression tree model
Description
E = cvloss(tree,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Alternatives
You can create a cross-validated tree model using crossval,
and call kfoldLoss instead of cvloss. If you are going to examine the cross-validated
tree more than once, then the alternative can save time.
However, unlike cvloss, kfoldLoss does not return SE,
Nleaf, or BestLevel.
Extended Capabilities
Version History
Introduced in R2011a
See Also
crossval | kfoldLoss | fitrtree | loss | RegressionTree