lassoPlot
Trace plot of lasso fit
Description
lassoPlot( creates a trace plot of the
values in B)B against the
L1 norm of B.
lassoPlot(
creates a plot with additional options specified by one or more name-value arguments.B,FitInfo,Name=Value)
Examples
Load the sample data.
load acetylenePrepare the design matrix for lasso fit with interactions. The x2fx function returns the quadratic model in the order of a constant term, linear terms and interaction terms: constant term, x1, x2, x3, x1.*x2, x1.*x3, and x2.*x3.
X = [x1 x2 x3]; D = x2fx(X,"interaction"); D(:,1) = []; % No constant term
Fit a regularized model of the data using the lasso function.
B = lasso(D,y);
Plot the lasso fits with labeled coefficients by using the PredictorNames name-value argument. Each line represents a trace of the values in B for a single predictor variable: x1, x2, x3, x1.*x2, x1.*x3, and x2.*x3.
lassoPlot(B,PredictorNames=["x1" "x2" "x3" "x1.*x2" "x1.*x3" "x2.*x3"]); legend(Location="NorthWest")

Load the sample data.
load acetylenePrepare the data for lasso fit with interactions.
X = [x1 x2 x3]; D = x2fx(X,"interaction"); D(:,1) = []; % No constant term
Fit a regularized model of the data using the lasso function.
[B,FitInfo] = lasso(D,y);
Plot the fits with the Lambda plot type and logarithmic scaling.
lassoPlot(B,FitInfo,PlotType="Lambda",XScale="log");
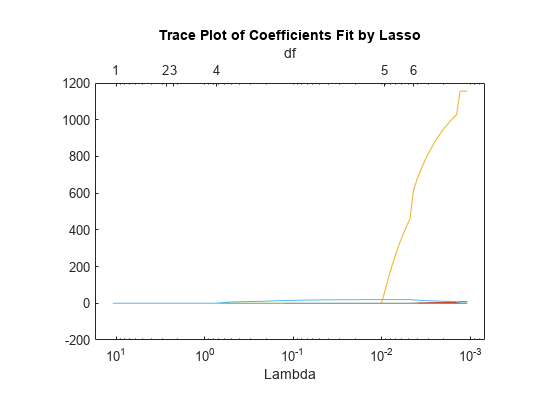
Visually examine the cross-validated error of various levels of regularization.
Load the sample data.
load acetyleneCreate a design matrix with interactions and no constant term.
X = [x1 x2 x3]; D = x2fx(X,"interaction"); D(:,1) = []; % No constant term
Construct the lasso fit using 10-fold cross-validation. Include the FitInfo output so you can plot the result.
rng default % For reproducibility [B,FitInfo] = lasso(D,y,CV=10);
Plot the cross-validated fits. The green circle and dotted line locate the Lambda with minimum cross-validation error. The blue circle and dotted line locate the point with minimum cross-validation error plus one standard error.
lassoPlot(B,FitInfo,PlotType="CV"); legend("show")

Input Arguments
Fit information of the generalized linear models, specified as a structure or a numeric vector.
When
FitInfois a structure, such as returned from thelassoorlassoglmfunction, thelassoPlotfunction creates a plot based on thePlotTypename-value argument.When
FitInfois a vector, thelassoPlotfunction forms the x-axis of the plot from the values inFitInfo. The length ofFitInfomust equal the number of columns ofB.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: lassoPlot(B,XScale="log") uses a logarithmic scaled
x-axis.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: lassoPlot(B,"XScale","log") uses a logarithmic scaled
x-axis.
Parent axes in which to draw the plot, specified as an Axes
object. If you do not specify Parent, then
lassoPlot creates the plot using the current axes. For more
information on creating an Axes object, see axes.
Plot type, specified as "L1", "Lambda", or
"CV". The PlotType argument applies when you
specify the FitInfo argument.
PlotType | Plot |
|---|---|
"L1" | lassoPlot creates the x-axis from
the L1 norm of the coefficients
in B. The x-axis at the top of the
plot contains the degrees of freedom (df), meaning the
number of nonzero coefficients of B. |
|
When
you choose this value, |
|
When you choose this value, |
If you include a cross-validated FitInfo structure,
lassoPlot also indicates two specific Lambda
values with green and blue dashed lines.
A green, dashed line indicates the value of
Lambdawith a minimum cross-validated mean squared error (MSE).A blue, dashed line indicates the greatest
Lambdathat is within one standard error of the minimum MSE. This Lambda value makes the sparsest model with relatively low MSE.
To display the label for each plot in the legend of the figure, type
legend("show") in the Command Window.
Predictor names, specified as a string array or cell array of character vectors.
The predictor names label each coefficient of B. If the length of
the PredictorNames argument is less than the number of rows of
B, the remaining labels are padded with default values.
If you do not specify this argument, but you specify the
FitInfo argument as a structure and the
PredictorNames field of the structure is not empty
({}), then the lassoPlot function uses those
predictor names instead of default values.
To display the legend with the predictor name as the label for each plot, type
legend("show") in the Command Window.
Scale of values along the x-axis, specified as one of these values.
"linear"— Linear x-axis. This is the default value when thePlotTypeargument is"L1"or"Lambda"."log"— Logarithmic scaled x-axis. This is the default value when thePlotTypeargument is"CV".
Data Types: string | char
Output Arguments
Axes of the plot, returned as an Axes object. For more information,
see Axes Appearance.
Figure window of the plot, returned as a Figure object. For more
information, see Special Object Identifiers.
Version History
Introduced in R2011b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)