medfreq
Median frequency
Syntax
Description
freq = medfreq(x)freq of the power
spectrum of a time-domain signal x. To compute the power
spectrum, medfreq uses the periodogram function with a
rectangular window and a number of DFT points equal to the length of
x.
freq = medfreq(___,freqRange)Fs
or f. If the second input is passed as empty,
medfreq assumes a normalized frequency. The default
value for freqRange is the entire bandwidth of the input
signal.
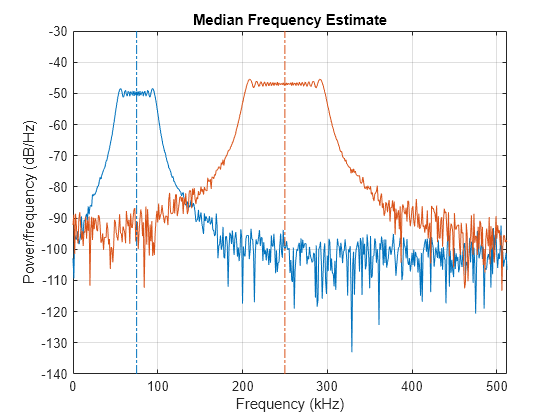
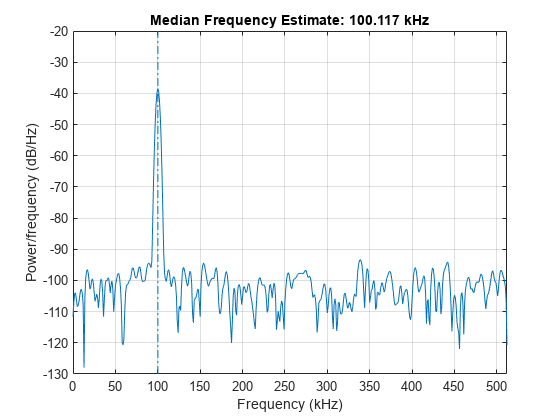
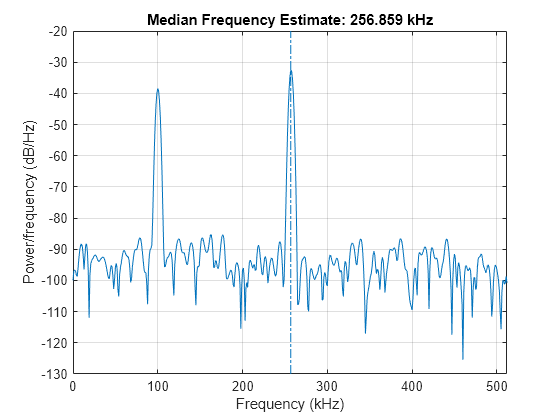
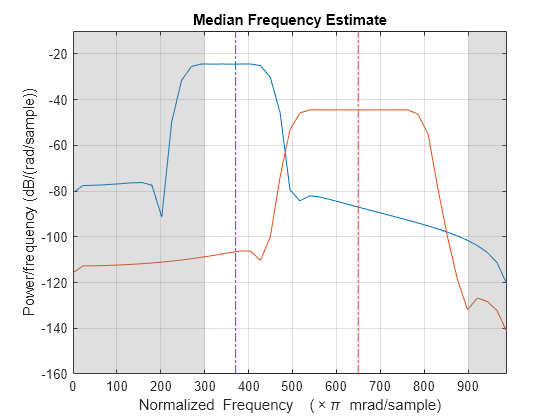
medfreq(___) with no output
arguments plots the PSD or power spectrum and annotates the median
frequency.
Examples
Input Arguments
Output Arguments
Algorithms
To determine the median frequency, medfreq computes a
periodogram power spectrum estimate using a rectangular window.
You can obtain the same value of median frequency, medFrq, from
a signal x at a sample rate Fs in these
three ways.
| Directly from the signal |
medFrq = medfreq(x,Fs) |
| From the periodogram of the signal |
[P,F] = periodogram(x,[],length(x),Fs); medFrq = medfreq(P,F) |
| From the power spectral estimate (Welch's PSD) of the signal |
[P,F] = pwelch(x,rectwin(length(x)),[],length(x),Fs); medFrq = medfreq(P,F) |
Note
Because medfreq uses an intermediary representation
to transform the input signal from the time domain to frequency domain, the
returned median frequency might vary, depending on the signal transformation
method, number of DFT points, and window size.
References
[1] Phinyomark, Angkoon, Sirinee Thongpanja, Huosheng Hu, Pornchai Phukpattaranont, and Chusak Limsakul. "The Usefulness of Mean and Median Frequencies in Electromyography Analysis." In Computational Intelligence in Electromyography Analysis – A Perspective on Current Applications and Future Challenges, edited by Ganesh R. Naik. London: IntechOpen, 2012. https://doi.org/10.5772/50639.
Extended Capabilities
Version History
Introduced in R2015aSee Also
findpeaks | meanfreq | periodogram | plomb | pwelch