imsegfmm
Binary image segmentation using fast marching method
Syntax
Description
[ also returns the normalized geodesic distance map
BW,D] =
imsegfmm(___)D computed using the fast marching method.
BW is a thresholded version of D,
where all the pixels that have normalized geodesic distance values less than or
equal to thresh are considered foreground pixels and set to
true. You can obtain different segmentation results by
thresholding D at different levels.
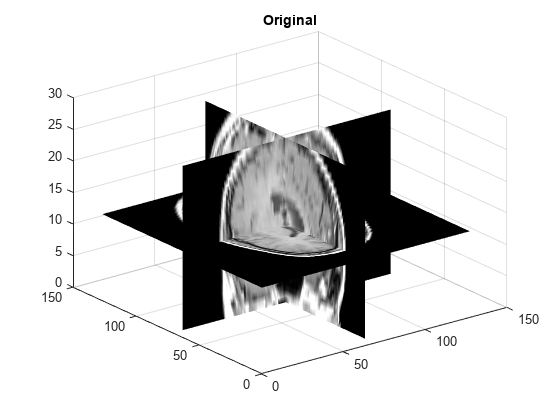
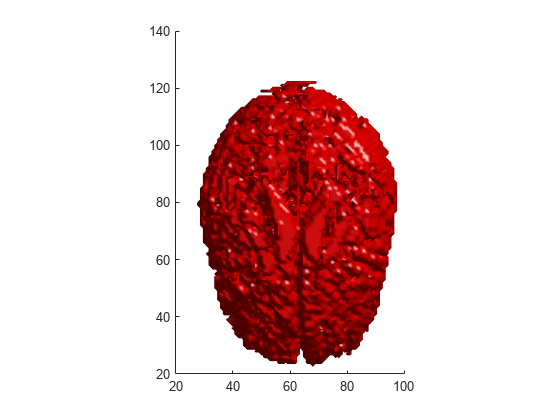
Examples
Input Arguments
Output Arguments
Tips
imsegfmmuses double-precision floating point operations for internal computations for all classes except classsingle. IfWis of data typesingle,imsegfmmuses single-precision floating point operations internally.imsegfmmsets pixels with0orNaNweight values toInfin the geodesic distance imageD. These pixels are part of the background in the segmented imageBW.
References
[1] Sethian, J. A. Level Set Methods and Fast Marching Methods: Evolving Interfaces in Computational Geometry, Fluid Mechanics, Computer Vision, and Materials Science, Cambridge University Press, 2nd Edition, 1999.
Extended Capabilities
Version History
Introduced in R2014bSee Also
activecontour | graydist | graydiffweight | gradientweight | Image Segmenter