Endmember Material Identification Using Spectral Library
This example shows how to identify the classes of endmember materials present in a hyperspectral image. The endmembers are pure spectral signature that signifies the reflectance characteristics of pixels belonging to a single surface material. The existing endmember extraction or identification algorithms extracts or identifies the pure pixels in a hyperspectral image. However, these techniques do not identify the material name or class to which the endmember spectrum belong to. In this example, you will extract the endmember signatures and then, classify or identify the class of an endmember material in the hyperspectral image by using spectral matching.
This example requires the Hyperspectral Imaging Library for Image Processing Toolbox™. You can install the Hyperspectral Imaging Library for Image Processing Toolbox from Add-On Explorer. For more information about installing add-ons, see Get and Manage Add-Ons. The Hyperspectral Imaging Library for Image Processing Toolbox requires desktop MATLAB®, as MATLAB® Online™ and MATLAB® Mobile™ do not support the library.
This example uses 1) the spectral signatures in the ECOSTRESS spectral library as the reference spectra and 2) a data sample from the Jasper Ridge dataset as the test data, for endmember material identification.
Read Reference Data from ECOSTRESS Spectral Library
Add the full file path containing the ECOSTRESS library files and specify the names of the files to be read from the library.
filenames = ["water.seawater.none.liquid.tir.seafoam.jhu.becknic.spectrum.txt",... "water.tapwater.none.liquid.all.tapwater.jhu.becknic.spectrum.txt",... "water.ice.none.solid.all.ice_dat_.jhu.becknic.spectrum.txt",... "vegetation.tree.eucalyptus.maculata.vswir.jpl087.jpl.asd.spectrum.txt",... "soil.utisol.hapludult.none.all.87p707.jhu.becknic.spectrum.txt",... "soil.mollisol.cryoboroll.none.all.85p4663.jhu.becknic.spectrum.txt",... "manmade.road.tar.solid.all.0099uuutar.jhu.becknic.spectrum.txt",... "manmade.concrete.pavingconcrete.solid.all.0092uuu_cnc.jhu.becknic.spectrum.txt"]; lib = readEcostressSig(filenames);
Display the lib data and inspect its values. The data is a struct variable specifying the class, subclass, wavelength, and reflectance related information.
lib
lib=1×8 struct array with fields:
Name
Type
Class
SubClass
ParticleSize
Genus
Species
SampleNo
Owner
WavelengthRange
Origin
CollectionDate
Description
Measurement
FirstColumn
SecondColumn
WavelengthUnit
DataUnit
FirstXValue
LastXValue
NumberOfXValues
AdditionalInformation
Wavelength
Reflectance
⋮
Plot the spectral signatures read from the ECOSTRESS spectral library.
figure hold on for idx = 1:numel(lib) plot(lib(idx).Wavelength,lib(idx).Reflectance,LineWidth=2); end axis tight box on xlabel("Wavelength (\mum)"); ylabel("Reflectance (%)"); classNames = {lib.Class}; legend(classNames,Location="northeast") title("Reference Spectra from ECOSTRESS Library"); hold off

Read Test Data
Read a test data from Jasper Ridge dataset by using the hypercube function. The function returns a hypercube object that stores the data cube and the metadata information read from the test data. The test data has 198 spectral bands and their wavelengths range from 399.4 nm to 2457 nm. The spectral resolution is up to 9.9 nm and the spatial resolution of each band image is 100-by-100. The test data contains four endmembers latent that includes road, soil, water, and trees.
hcube = imhypercube("jasperRidge2_R198.hdr");Extract Endmember Spectra
To compute the total number of spectrally distinct endmembers present in the test data, use the countEndmembersHFC function. This function finds the number of endmembers by using the Harsanyi–Farrand–Chang (HFC) method. Set the probability of false alarm (PFA) to a low value in order to avoid false detections.
numEndmembers = countEndmembersHFC(hcube,PFA=10^-27);
Extract the endmembers of the test data by using the N-FINDR method.
endMembers = nfindr(hcube,numEndmembers);
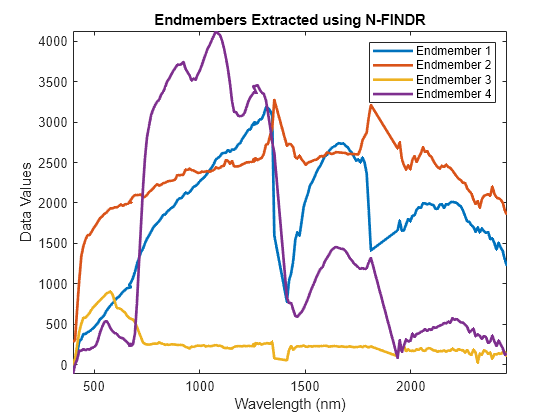
Read the wavelength values from the hypercube object hcube. Plot the extracted endmember signatures. The test data comprises of 4 endmember materials and the class names of these materials can be identified through spectral matching.
figure plot(hcube.Wavelength,endMembers,LineWidth=2) axis tight xlabel("Wavelength (nm)") ylabel("Data Values") title("Endmembers Extracted using N-FINDR") num = 1:numEndmembers; legendName = "Endmember "+num; legend(legendName)
Identify Endmember Material
To identify the name of an endmember material, use the spectralMatch function. The function computes the spectral similarity between the library files and an endmember spectrum to be classified. Select spectral information divergence (SID) method for computing the matching score. Typically, a low value of SID score means better matching between the test and the reference spectra. Then, the test spectrum is classified to belong to the class of the best matching reference spectrum.
For example, to identify the class of the third and fourth endmember material, find the spectral similarity between the library signatures and the respective endmember spectrum. The index of the minimum SID score value specifies the class name in the spectral library. The third endmember spectrum is identified as Sea Water and the fourth endmember spectrum is identified as Tree.
wavelength = hcube.Wavelength; detection = cell(1,1); cnt = 1; queryEndmember = [3 4]; for num = 1:numel(queryEndmember) spectra = endMembers(:,queryEndmember(num)); scoreValues = spectralMatch(lib,spectra,wavelength,Method="sid"); [~, matchIdx] = min(scoreValues); detection{cnt} = lib(matchIdx).Class; disp("Endmember spectrum "+queryEndmember(num)+" is identified as "+detection{cnt}) cnt=cnt+1; end
Endmember spectrum 3 is identified as Sea Water Endmember spectrum 4 is identified as Tree
Segment Endmember Regions in Test Data
To visually inspect the identification results, localise and segment the image regions specific to the endmember materials in the test data. Use the sid function to compute pixel-wise spectral similarity between the pixel spectrum and the extracted endmember spectrum. Then, perform thresholding to segment the desired endmember regions in the test data and generate the segmented image. Select the value for threshold as 15 to select the best matching pixels.
For visualization, create an RGB version of the test data by using the colorize function. Create an overlay of the segmented Sea Water and Tree endmember regions over the RGB image.
threshold = 15; rgbImg = colorize(hcube,Method="rgb",ContrastStretching=true); overlayImg = rgbImg; labelColor = ["Blue","Green"]; datacube = gather(hcube); segmentedImg = cell(size(datacube,1),size(datacube,2),numel(queryEndmember)); for num = 1:numel(queryEndmember) scoreMap = sid(hcube,endMembers(:,queryEndmember(num))); segmentedImg{num} = scoreMap <= threshold; overlayImg = imoverlay(overlayImg,segmentedImg{num},labelColor{num}); end
Display Results
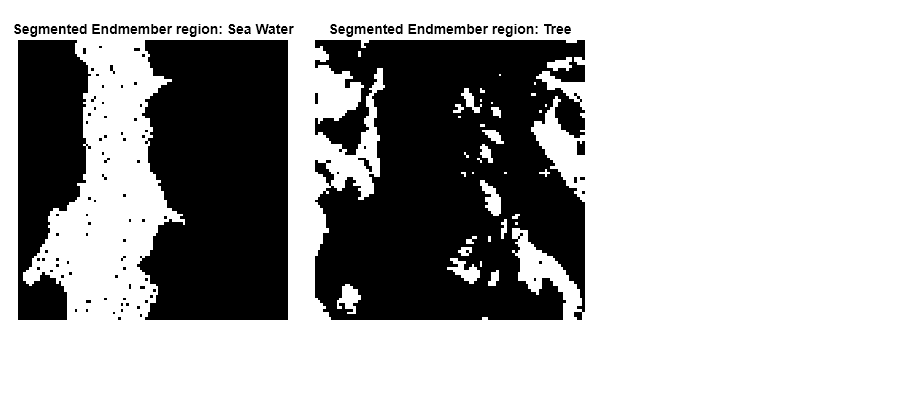
Visually inspect the identification results. Display the binary segmentation results of the Sea Water and Tree endmember regions.
figure(Position=[0 0 900 400]) plotdim = [0.02 0.2 0.3 0.7;0.35 0.2 0.3 0.7]; for num = 1:numel(queryEndmember) subplot(Position=plotdim(num,:)) imagesc(segmentedImg{num}) title("Segmented Endmember region: "+detection{num}); colormap([0 0 0;1 1 1]) axis off end
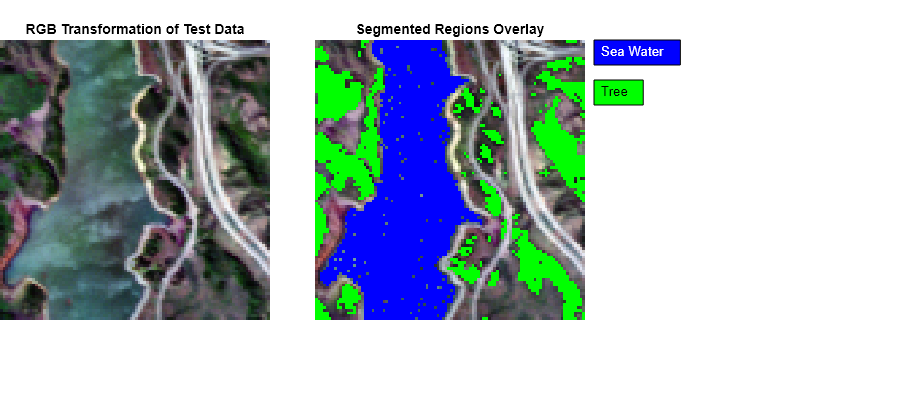
Display the overlay the endmember regions over the RGB image.
figure(Position=[0 0 900 400]) subplot(Position=[0 0.2 0.3 0.7]) imagesc(rgbImg) title("RGB Transformation of Test Data"); axis off subplot(Position=[0.35 0.2 0.3 0.7]) imagesc(overlayImg) title("Segmented Regions Overlay") hold on dim = [0.66 0.6 0.3 0.3]; annotation("textbox",dim,String="Sea Water",Color=[1 1 1],BackgroundColor=[0 0 1],FitBoxToText="on"); dim = [0.66 0.5 0.3 0.3]; annotation("textbox",dim,String="Tree",BackgroundColor=[0 1 0],FitBoxToText="on"); hold off axis off
References
[1] Kruse, F.A., A.B. Lefkoff, J.W. Boardman, K.B. Heidebrecht, A.T. Shapiro, P.J. Barloon, and A.F.H. Goetz. “The Spectral Image Processing System (SIPS)—Interactive Visualization and Analysis of Imaging Spectrometer Data.” Remote Sensing of Environment 44, no. 2–3 (May 1993): 145–63. https://doi.org/10.1016/0034-4257(93)90013-N.
[2] ECOSTRESS Spectral Library: https://speclib.jpl.nasa.gov
[3] Meerdink, Susan K., Simon J. Hook, Dar A. Roberts, and Elsa A. Abbott. “The ECOSTRESS Spectral Library Version 1.0.” Remote Sensing of Environment 230 (September 2019): 111196. https://doi.org/10.1016/j.rse.2019.05.015.
[4] Baldridge, A.M., S.J. Hook, C.I. Grove, and G. Rivera. “The ASTER Spectral Library Version 2.0.” Remote Sensing of Environment 113, no. 4 (April 2009): 711–15. https://doi.org/10.1016/j.rse.2008.11.007.
See Also
hypercube | colorize | spectralMatch | readEcostressSig | countEndmembersHFC