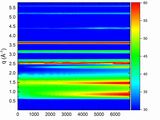
time-resolved WAXS - cutting q-cuts as a function of phi
Version 1.0.0 (3.62 KB) by
Karol Vegso
This program performs time-binning and pixel-binning on WAXS images. It cuts q-cuts as a function of azimuthal angle phi.
This m-file or script uses GIXSDATA from Argonne National Laboratory / Advanced Photon source. It was written by Zhang Jiang. Please download latest version of GIXSGUI on the web-page <https://www.aps.anl.gov/Sector-8/8-ID/Operations-and-Schedules/Useful-Links/Sector-8-GIXSGUI>. Then, unzip GIXSGUI folder. Start Matlab. On the Home tab, click "Set Path" > "Add with Subfolders..." and select the folder that contains the GIXSGUI distribution. Click "Save" and "Close".
Then, open my script "tb_pb_rot_WAXS_q_cuts_as_func_phi.m". In the name of m-file, tb means time-binning, pb means pixel-binning, rot means that rotation of images is possible, and q-cuts means that program extract q-cuts as a function of azimuthal angle phi.
In program, you need to specify the path to folder with your images. Therefore, you need to fill string variable path_to_input_folder (e.g. path_to_input_folder='d:\Pilatus_images\';).
You need to enter path to output folder, where your results will be saved. Therefore, you need to fill string variable path_to_output_folder (e.g. path_to_output_folder='d:\output_folder\';).
You need to enter path to your text calibration file, where your calibration parameters are stored. Therefore, you need to fill string variable path_to_calib_file (e.g. path_to_calib_file='d:\calibration\calibration_text_file.txt';).
To create calibration text file, please create text file somewhere on your computer. Then, fill one row in text file for example like:
Pilatus 487 407 0.172 0.172 8.04 510 1 1 4 0.5 0.0 263 383 263 383
1-st string is type of your camera, it can be for example Pilatus from Decrtris. If not, there can be written Other.
The 2-nd unsigned integer is number of pixels in horizontal direction and 3-rd unsigned integer is number of pixels in vertical direction. For example, here, I use Pilatus 200K with area of 487 x 407 pixels (H X V).
The 4-th float number is size of pixel in horizontal direction and 5-th float number is size of pixel in vertical direction. Here, I use Pilatus detector, so, square pixel size is 0.172 x 0.172 mm (H x V).
The 6-th float number is energy of X-rays in keV. Here, I use laboratory microfocus X-ray source with Cu anode. The energy of CuKalpha radiation in this example is 8.04 keV.
The 7-th float number is sample to detector distance, also called SDD in mm.
The 8-th unsigned integer is geometry in GIXSDATA or GIXSGUI. There are only two possibilities for geometry value. It can be 1 for transmission or 2 for reflection. Of course, here, we evaluate WAXS data in transmission mode. It means geometry must be 1. We are in transmission mode.
The 9-th unsigned integer is for phi mode value. According to the GIXSDATA and GIXSGUI manual, Range for pixel azimuthal angle with respect to the direct beam. [1]/2/3/4 for [(−180° , 180°)]/(0°, 360°)/(−270°, 90°)/(−90°, 270°). It is OK, to put here number 1 for azimuthal range (−180°, 180°). Then, if you want to integrate full azimuthal range (entire image), you will integrate from phi=-180° to phi=+180°. You can also set azimuthal range to value 2, it means that your azimuthal range will be (0°, 360°). Then, if you want to integrate full azimthal range (entire image), you will integrate from phi=0° to phi=+360°.
The 10-th unsigned integer is polarization mode. According to the GIXSDATA and GIXSGUI manual, Incident beam polarization. 1/[2]/3/4 for none/[horizontal] /vertical/unpolarized. In my opinion, if your data were measured at synchrotron beamline, you need to set e.g. number 2 for horizontal polarization. If your data were measured with laboratory microfocus X-ray source, you need to set e.g. number 4 for unpolarized beam.
The 11-th float number is Horizontal Polarization Fraction. It can be set only for horizontal polarization mode (polarization mode = number 2) or vertical polarization mode (polarization mode = number 3). If Horizontal Polarization Fraction = 0.5, you have half of horizontal polarization and half of vertical polarization. For unpolarized beam, the Horizontal Polarization Fraction is simply not used.
The 12-th float number is angle of incidence measured with respect to the sample surface. I put there value 0.0, because for transmission mode, this number is irrelevant. You are not in grazing-incidence geometry, but in transmission.
The 13-th float number is position of primary beam in the image in horizontal direction. The 14-th float number is position of primary beam in the image in vertical direction. Here, in this example, Beam0_Hor = 263 pixel, and Beam0_Ver = 383 pixel.
The 15-th float number is position of specular/reflected beam in the image in horizontal direction. Here, I put some dummy numbers like Specular_Hor = 263 pixel, and Specular_Ver = 383 pixel. For transmission mode, this number is irrelevant. You are not in grazing-incidence geometry, but in transmission.
In my script, calibration text file is read by readtable function. The calibration in the script is between 60 and 92 lines. Read it carefully to understand calibration.
If you perform time-resolved study with Pilatus detector, your images are like
image_00000.tif
image_00001.tif
image_00002.tif
image_00003.tif
and so on until
image_01199.tif
image_01200.tif
Here, your first image is image_00000.tif and last image is image_01200.tif.
According to this example, your image name root is image_name_root=''image".
Here for Pilatus detector, the numbering contains 5 digits for example 10-th image has numbering 00009. Therfore, image_no_width=5;
The underscore between image_name_root and image numbering is added automatically.
You also need to fill image name extension. It can be '.tif' or '.cbf' (crystallographic binary file). You need to fill string variable for example image_name_ext='.tif';
If you performed time resolved study, where you have 1201 Pilatus tif images, then your first image is image_00000.tif and your last image is image_01200.tif. Then, you fill no_image_start=0 and no_image_stop=1200;
This software can perform time binning. Therefore, you need to fill unsigned integer variable time_binning.
If you don't want to use time binning, then time_binning = 1;
If you want to sum each two images, then time_binning = 2;
But be carefull, for time_binning = 1, it does not affect no_image_start and no_image_stop variables.
But for time_binning = 2, no_image_start=0 and no_image_stop=1199, beacuse from 0 to 1199 image, we have 1200 images which gives remainder 0 after division by 2. If no_image_start=0 and no_image_stop=1200, then you have 1200 +1 = 1201 images, which gives remainder 1 after disvison by 2. Then, program would crash.
My script uses imrotate function from Image Processing Toolbox, so, you can rotate images if necessary. You can fill, variable rot_angle in degrees. For many of your applications, rot_angle=0.0; % [deg].
In my script, you can perform pixel binning in horizontal direction (x direction) and vertical direction (y direction).
The horizontal or x direction is given by number of columns in image.
The vertical or y direction is given by number of rows in image.
For no pixel binning in horizontal or x direction:
% specify pixel binning in column direction
pixel_binning_col=1;
% start pixel in column direction
start_pixel_col=1; % [pixels]
% stop pixel in col direction
stop_pixel_col=487; % [pixels]
This is for eaxmple for Pialtus 200K detector, which has 487 pixels in horizontal or x direction (or 487 columns in image matrix). For no pixel binning, you go from 1-st pixel (start_pixel_col) to 487-th pixel (stop_pixel_col).
For no pixel binning in vertical or y direction:
% specify pixel binning in row direction
pixel_binning_row=1;
% start pixel in row direction
start_pixel_row=1; % [pixels]
% stop pixel in row direction
stop_pixel_row=407; % [pixels]
This is for eaxmple for Pialtus 200K detector, which has 407 pixels in vertical or y direction (or 407 rows in image matrix). For no pixel binning, you go from 1-st pixel (start_pixel_row) to 407-th pixel (stop_pixel_row).
Now consider that you want to sum each 2 x 2 pixels (H x V). So, you want to perform 2 by 2 pixel binning. Then, you set, pixel_binning_col=2; pixel_binning_row=2;
The setting in script should be like that:
% specify pixel binning in column direction
pixel_binning_col=2;
% start pixel in column direction
start_pixel_col=1; % [pixels]
% stop pixel in col direction
stop_pixel_col=486; % [pixels]
and
% specify pixel binning in row direction
pixel_binning_row=2;
% start pixel in row direction
start_pixel_row=1; % [pixels]
% stop pixel in row direction
stop_pixel_row=406; % [pixels]
So, what was changed? I changed stop_pixel_col to value 486, because from start_pixel_col=1 to stop_pixel_col=486, you have 486 pixels which gives remainder 0 after division by 2 (pixel_binning_col=2).
I changed stop_pixel_row to value 406, because from start_pixel_row=1 to stop_pixel_row=406, you have 406 pixels which gives remainder 0 after division by 2 (pixel_binning_row=2).
Now, you can consider that you want pixel binning 4x4.
Then
% specify pixel binning in column direction
pixel_binning_col=4;
% start pixel in column direction
start_pixel_col=1; % [pixels]
% stop pixel in col direction
stop_pixel_col=484; % [pixels] (remainder 0 after disvison by 4)
and
% specify pixel binning in row direction
pixel_binning_row=4;
% start pixel in row direction
start_pixel_row=1; % [pixels]
% stop pixel in row direction
stop_pixel_row=404; % [pixels] (remainder 0 after disvison by 4)
The q-cuts parameter or linecut parameter are between lines 92 and 107. Here, you cut from q_min to q_max value in reciprocal Angstroms. The integration of q-cuts happens between phi_min and phi_max values. The phi angle is azimuthal angle of the reciprocal space in WAXS geometry or transmission mode. You need to specify how many points you need to cut along q direction from q_min to q_max. Therefore, you must specify no_of_q_points variable. If no_of_q_points will be too big, you can oversample your data / linecut.
The q-cut or linecut is performed in the lines 225 and 234. The q-cut is intensity plot where X-variable contains q values and Y-variable are intensity values. The X-variable is set by xflag value equal to 1 (q, QMap). For details see page 20 of GIXSGUI / GIXSDATA manual, where xflag is specified.
The constraint is set in the line 228 like constr = [1 2 phi_min phi_max; 1 1 q_min q_max];. You can read it as (1 2 phi_min phi_max) = (AND (constraint put on phi variable (flag=2)) integrate from (minimal value of phi) to (maximal value of phi)). You can read it as (1 1 q_min q_max) = (AND (constraint put on q variable (flag=1)) integrate from (minimal value of q) to (maximal value of q)).
The time-binning of GIWAXS images happens between 118 and 166 lines. The time-binned image is stored in matrix integrate_image. The pixel-binned image is stored in matrix new_image. It happnes between 166 and 181 lines.
The new gixsdata object is created in line 209. The time-binned and pixel-binned image represented by matrix new_image is stored into the new gixsdata object (obj.RawData=new_image(:,:);).
Your final q-cuts in time are stored in the matrix buffer_linecut and your q axis is stored in the x vector. The q-cuts matrix represented by buffer_linecut and q axis represented by x vector are saved as text matrices in the output folder.
Note for programmers: The GIXSDATA and GIXSGUI is written as object oriented program. Therefore, you need to create gixsdata object, e.g. please see line 139 with obj=gixsdata(full_path_to_image);. It creates object for existing time-resolved image in time-binning loop. The object is destroyed or deleted in line 163. The time-binned image is pixel-binned between lines 166 a6nd 181. Then, new gixsdata object is created in line 209 (obj=gixsdata(full_path_to_image_create);) for time-binned and pixel-binned image. The pixel-binned image is calibrated and reshaped to reciprocal space. The object for time-binned and pixel-binned image is destroyed or deleted in line 235.
Cite As
Karol Vegso (2024). time-resolved WAXS - cutting q-cuts as a function of phi (https://www.mathworks.com/matlabcentral/fileexchange/113385-time-resolved-waxs-cutting-q-cuts-as-a-function-of-phi), MATLAB Central File Exchange. Retrieved .
MATLAB Release Compatibility
Created with
R2021b
Compatible with R2021b to R2022a
Platform Compatibility
Windows macOS LinuxTags
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.
| Version | Published | Release Notes | |
|---|---|---|---|
| 1.0.0 |