Specify Starting Points and Values for surrogateopt, Problem-Based
For some solvers, you can pass the objective and constraint function values, if any, to solve in the x0 argument. This can save time in the solver. Pass a vector of OptimizationValues objects. Create this vector using the optimvalues function.
The solvers that can use the objective function values are:
gagamultiobjparetosearchsurrogateopt
The solvers that can use nonlinear constraint function values are:
paretosearchsurrogateopt
For example, minimize the peaks function using surrogateopt, starting with values from a grid of initial points. Create a grid from -10 to 10 in the x variable, and –5/2 to 5/2 in the y variable with spacing 1/2. Compute the objective function values at the initial points.
x = optimvar("x",LowerBound=-10,UpperBound=10); y = optimvar("y",LowerBound=-5/2,UpperBound=5/2); prob = optimproblem(Objective=peaks(x,y)); xval = -10:10; yval = (-5:5)/2; [x0x,x0y] = meshgrid(xval,yval); peaksvals = peaks(x0x,x0y);
Pass the values in the x0 argument by using optimvalues. This saves time for solve, as solve does not need to compute the values. Pass the values as row vectors.
x0 = optimvalues(prob,x=x0x(:)',y=x0y(:)',...
Objective=peaksvals(:)');Solve the problem using surrogateopt with the initial values.
[sol,fval,eflag,output] = solve(prob,x0,Solver="surrogateopt")Solving problem using surrogateopt.
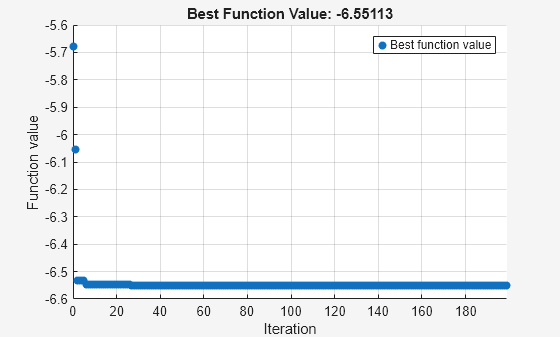
surrogateopt stopped because it exceeded the function evaluation limit set by 'options.MaxFunctionEvaluations'.
sol = struct with fields:
x: 0.2279
y: -1.6258
fval = -6.5511
eflag =
SolverLimitExceeded
output = struct with fields:
elapsedtime: 35.9581
funccount: 200
constrviolation: 0
ineq: [1×1 struct]
rngstate: [1×1 struct]
message: 'surrogateopt stopped because it exceeded the function evaluation limit set by ↵'options.MaxFunctionEvaluations'.'
solver: 'surrogateopt'
See Also
surrogateopt | solve | optimvalues